Monte Carlo Simulations
What is Monte Carlo Simulations?
One of the main motivations to switch from spreadsheet-type tools (such
as Microsoft Excel) to a program like R is for simulation modeling. R
allows us to repeat the same (potentially complex and detailed)
calculations with different random values over and over again.
Within the same software, we can then summarize and plot the results of
these replicated calculations. Monte Carlo methods are used to perform
this type of analysis they randomly sample from a set of values in order
to generate and summarize a distribution of some statistic related to
the sampled quantities.
Randomness
Random processes are an important aspect of simulation modeling. A
random process is one that produces a different result each time it is
run according to some rules. They are inextricably tied to the concept
of uncertainty, you have no idea what will happen the next time the
process is run.
There are two basic ways to introduce randomness in R
Random deviattes
Resampling
Random Deviates
Each person alive at the start of the year has the option of living or
dying at the conclusion of the year. There are two possible endings
here, and each person has an 80% probability of surviving. survive is
the outcome of a binomial random process in which there were n
individuals alive at the start of this year and p is the probability
that any one of them would live to the next year.
In R, we can simulate a binomial random process with p=0.8 and n=100.
rbinom(n= 1, size =100,
prob= 0.8)## [1] 80At this time I got 73, but we almost certainly get different number than
this one.
With a little tinkering, we can also plot it
survivors = rbinom(1000,
100, 0.8)
hist(survivors,
col = "skyblue")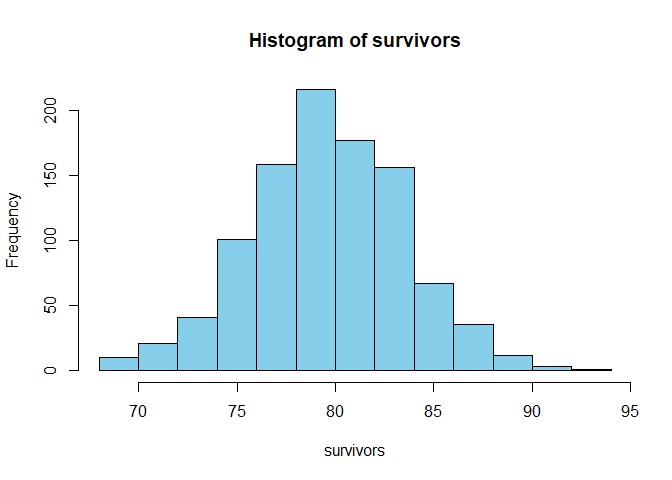
We could also used other processes like log normal
The log normal process is another random process. It creates random
numbers using a log of the values that is regularly distributed, with a
mean of log mean and a standard deviation of log sd.
hist(rlnorm(1000,0,0.1),col="skyblue")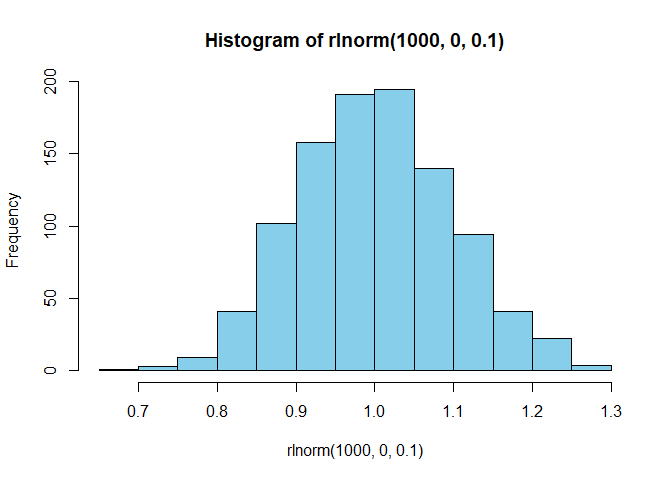
Need for sampling
There are several situations in probability, and more broadly in machine
learning, where an analytical solution cannot be calculated immediately.
In Machine Learning, a problem of class imbalance exists. In fact, some
would argue that for most practical probabilistic models, accurate
inference is impossible.
The desired calculation is usually a sum of discrete distributions or an
integral of continuous distributions, and thus is computationally
difficult. For a variety of reasons, such as the huge number of random
variables, the stochastic nature of the domain, noise in the data, a
shortage of observations, and more, the calculation may be intractable.
Resampling
Using random deviates to generate fresh random numbers is excellent, but
what if we already have a set of numbers to which we want to add
randomness? We can utilize resampling techniques to do this. To sample
size elements from the vector x in R, use the sample() function.
Resampling of 1 to 10
sample(x = 1:10, size =5)## [1] 4 3 10 9 2Sample with replacement
sample(x = c("a","b","c"), size = 10, replace = T)## [1] "a" "a" "a" "b" "a" "b" "a" "c" "b" "b"Sample with set probalilities
sample(x = c("live","die"),size = 10, replace = T, prob = c(0.8,0.2))## [1] "live" "live" "die" "die" "live" "live" "live" "die" "live" "live"Reproducing Randomness
We may want to receive the same precise random integers each time we run
our script for reproducibility. To do so, we must first set the random
seed, which is the starting point of our computer’s random number
generator.
set.seed(1234)
rnorm(1)## [1] -1.207066Let’s try without random seed
rnorm(1)## [1] 0.2774292Each time we get different result.
Replication
To use Monte Carlo methods, we need to be able to replicate some random
process many times. There are two main ways this is commonly done,
either with replicate()or with for()loops.
The replicate() functions executes same expression many times and
returns the output from each execution. Say we have a vector x, which
represents 40 observations of an animal length(mm).
x = rnorm(30, 500,40)We want to create the mean length sampling distribution “by hand.” We
can take a random sample, determine the mean, and then repeat the
process as many times as necessary.
Replication with “for” loop
A loop is a command in programming that repeats itself until it reaches
a specified point. R has a few types of loops, repeat(), while(), and
for(). for() loops are among the most common in simuation modeling.For
each value in a vector, a for() loop performs an operation for the
number of times you specify.
For loop syntax
for(var in seq){ expression(var) }
for( i in 1:5){
print(i^2)
}## [1] 1
## [1] 4
## [1] 9
## [1] 16
## [1] 25nt = 100
N= NULL
N[1] = 1000
for(t in 2:nt){
N[t] = (N[t-1]*1.1*rlnorm(1,0,0.1))*(1-0.08)
}Let’s plot it
plot(N, type= "l", pch = 15, xlab = "Year", ylab = "Abundance")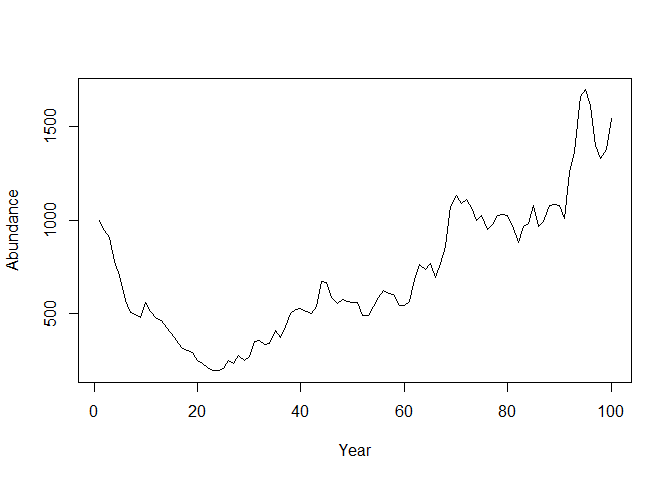
Summarization of simulation
After replicating a calculation many times, we will need to summarize
the results.
Simulating Based Learning
mu = 500
sig = 30
random = rnorm(100,mu,sig)
p = seq(0.01, 0.99, 0.01)
random_q = quantile(random,p)
normal_q = qnorm(p,mu,sig)
plot(normal_q~random_q)
abline(c(0,1))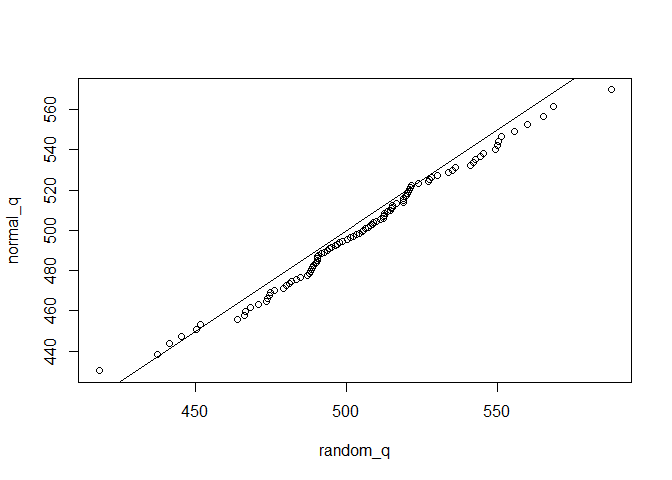
q = seq(400,600,10)
random_cdf = ecdf(random)
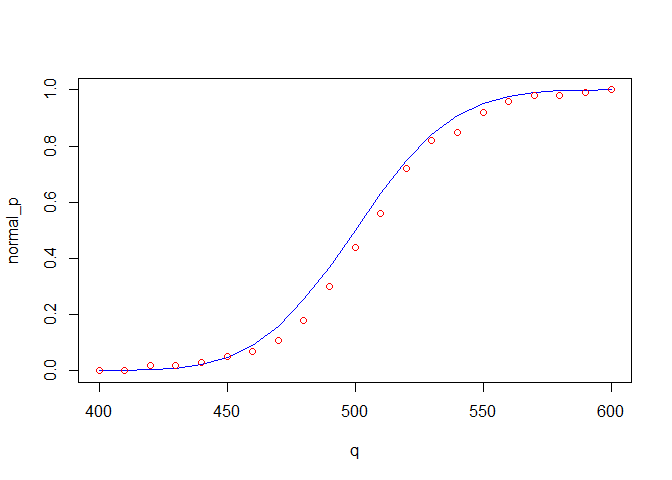
random_p =random_cdf(q)
normal_p = pnorm(q,mu,sig)
plot(normal_p~q, type= "l", col = "blue")
points(random_p~q,col = "red")