Check the data mtcars with head and save a new data as mtcars.subset after dropping two non-numeric (binary) variables for PCA analysis
data <- mtcars
head(data)## mpg cyl disp hp drat wt qsec vs am gear carb
## Mazda RX4 21.0 6 160 110 3.90 2.620 16.46 0 1 4 4
## Mazda RX4 Wag 21.0 6 160 110 3.90 2.875 17.02 0 1 4 4
## Datsun 710 22.8 4 108 93 3.85 2.320 18.61 1 1 4 1
## Hornet 4 Drive 21.4 6 258 110 3.08 3.215 19.44 1 0 3 1
## Hornet Sportabout 18.7 8 360 175 3.15 3.440 17.02 0 0 3 2
## Valiant 18.1 6 225 105 2.76 3.460 20.22 1 0 3 1str(data)## 'data.frame': 32 obs. of 11 variables:
## $ mpg : num 21 21 22.8 21.4 18.7 18.1 14.3 24.4 22.8 19.2 ...
## $ cyl : num 6 6 4 6 8 6 8 4 4 6 ...
## $ disp: num 160 160 108 258 360 ...
## $ hp : num 110 110 93 110 175 105 245 62 95 123 ...
## $ drat: num 3.9 3.9 3.85 3.08 3.15 2.76 3.21 3.69 3.92 3.92 ...
## $ wt : num 2.62 2.88 2.32 3.21 3.44 ...
## $ qsec: num 16.5 17 18.6 19.4 17 ...
## $ vs : num 0 0 1 1 0 1 0 1 1 1 ...
## $ am : num 1 1 1 0 0 0 0 0 0 0 ...
## $ gear: num 4 4 4 3 3 3 3 4 4 4 ...
## $ carb: num 4 4 1 1 2 1 4 2 2 4 ...In our data vs and am are binary variable so I drop them here.
library(dplyr)
mtcars.subset <- data[,-c(8,9)]Fit PCA in the data by mtcars.pca matcars.subset with cor = TRUE and scores = TRUE
mtcars.pca<-prcomp(mtcars.subset, cor = TRUE, scores = TRUE)## Warning: In prcomp.default(mtcars.subset, cor = TRUE, scores = TRUE) :
## extra arguments 'cor', 'scores' will be disregardedGet summary of mtcars.pca
summary(mtcars.pca)## Importance of components:
## PC1 PC2 PC3 PC4 PC5 PC6 PC7
## Standard deviation 136.532 38.14735 3.06642 1.27492 0.90474 0.64734 0.3054
## Proportion of Variance 0.927 0.07237 0.00047 0.00008 0.00004 0.00002 0.0000
## Cumulative Proportion 0.927 0.99938 0.99985 0.99993 0.99997 0.99999 1.0000
## PC8 PC9
## Standard deviation 0.2859 0.2159
## Proportion of Variance 0.0000 0.0000
## Cumulative Proportion 1.0000 1.0000From above summary we see when standard deviation is greater propertion
of variance is high similarly when standard deviation is low propertion
of variation is also low.
Get eigenvalue of the components using standard deviation of mtcars.pca and chose the number of components based on Kaiser’s criteria
mtcars.pca$sdev ^2## [1] 1.864106e+04 1.455220e+03 9.402948e+00 1.625431e+00 8.185525e-01
## [6] 4.190430e-01 9.327903e-02 8.175127e-02 4.660443e-02Get scree plot and chose the number of components best on “first bend” of this plot
#Calculating total variance explained by each principal component
var_explained<-mtcars.pca$sdev^2/sum(mtcars.pca$sdev^2)
##Creating scree plot
library(ggplot2)
qplot(c(1:9), var_explained) + geom_line() + xlab("Principal Component") + ylab("Variance explained") + ggtitle("Scree Plot")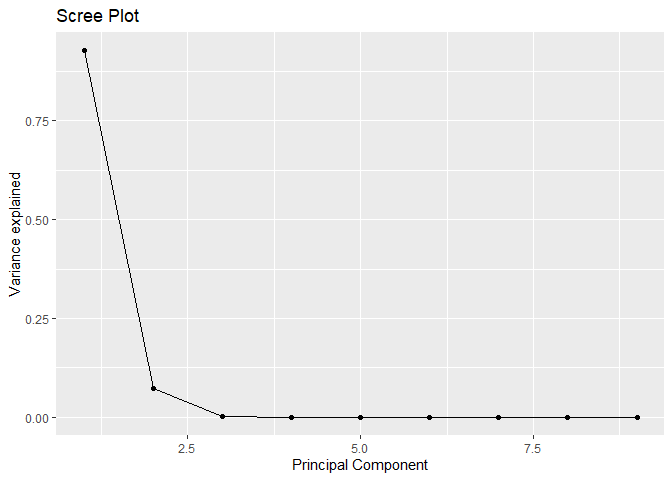
How many components must be retained based on Kaiser’s rule and/or scree plot?
Kisher’s rule suggaest us to use 2 components and Scree plot
suggest us to retain 4 components for the problem.
Fit the final PCA model based on the retained components
library(psych)## Warning: package 'psych' was built under R version 4.1.2mtcars.pca<- psych::principal(mtcars.subset, nfactors = 4, rotate = "none")
mtcars.pca## Principal Components Analysis
## Call: psych::principal(r = mtcars.subset, nfactors = 4, rotate = "none")
## Standardized loadings (pattern matrix) based upon correlation matrix
## PC1 PC2 PC3 PC4 h2 u2 com
## mpg -0.93 0.04 -0.16 0.00 0.90 0.0995 1.1
## cyl 0.96 0.02 -0.18 0.02 0.95 0.0504 1.1
## disp 0.94 -0.13 -0.06 0.17 0.94 0.0569 1.1
## hp 0.87 0.39 -0.01 0.04 0.91 0.0854 1.4
## drat -0.74 0.49 0.11 0.44 0.99 0.0062 2.5
## wt 0.89 -0.25 0.32 0.10 0.96 0.0360 1.5
## qsec -0.53 -0.70 0.45 -0.02 0.97 0.0283 2.6
## gear -0.50 0.79 0.15 -0.15 0.92 0.0775 1.8
## carb 0.58 0.70 0.33 -0.11 0.95 0.0525 2.5
##
## PC1 PC2 PC3 PC4
## SS loadings 5.66 2.08 0.50 0.27
## Proportion Var 0.63 0.23 0.06 0.03
## Cumulative Var 0.63 0.86 0.92 0.95
## Proportion Explained 0.66 0.24 0.06 0.03
## Cumulative Proportion 0.66 0.91 0.97 1.00
##
## Mean item complexity = 1.7
## Test of the hypothesis that 4 components are sufficient.
##
## The root mean square of the residuals (RMSR) is 0.02
## with the empirical chi square 0.96 with prob < 0.99
##
## Fit based upon off diagonal values = 1Get the head of the saved loadings of mtcars.pca
head(mtcars.pca)## $values
## [1] 5.65593947 2.08210029 0.50421482 0.26502753 0.18315864 0.12379319 0.10506192
## [8] 0.05851375 0.02219038
##
## $rotation
## [1] "none"
##
## $n.obs
## [1] 32
##
## $communality
## mpg cyl disp hp drat wt qsec gear
## 0.9004692 0.9495949 0.9430951 0.9146434 0.9938301 0.9639755 0.9716949 0.9224616
## carb
## 0.9475174
##
## $loadings
##
## Loadings:
## PC1 PC2 PC3 PC4
## mpg -0.935 -0.157
## cyl 0.957 -0.179
## disp 0.945 -0.128 0.175
## hp 0.873 0.389
## drat -0.742 0.493 0.106 0.435
## wt 0.888 -0.248 0.322
## qsec -0.534 -0.698 0.446
## gear -0.498 0.795 0.147 -0.145
## carb 0.582 0.699 0.330 -0.110
##
## PC1 PC2 PC3 PC4
## SS loadings 5.656 2.082 0.504 0.265
## Proportion Var 0.628 0.231 0.056 0.029
## Cumulative Var 0.628 0.860 0.916 0.945
##
## $fit
## [1] 0.9982615Retain two components, get their loadings
mtcars.pca_2 <- principal(mtcars.subset, nfactors = 2, rotate="none")
mtcars.pca_2$loadings##
## Loadings:
## PC1 PC2
## mpg -0.935
## cyl 0.957
## disp 0.945 -0.128
## hp 0.873 0.389
## drat -0.742 0.493
## wt 0.888 -0.248
## qsec -0.534 -0.698
## gear -0.498 0.795
## carb 0.582 0.699
##
## PC1 PC2
## SS loadings 5.656 2.082
## Proportion Var 0.628 0.231
## Cumulative Var 0.628 0.860Principal components (PCs) are constructed by the linear combination of
the original variables, where PCA loading are the coefficients. Here,
cyl has the weights of 0.957 on PC1 computation but not in PC2. Positive
loading in above data indicates a variable and a component are
positively correlated. Negative loading indicate a negative correlation
between the variable and component. Similarly disp has positive loading
with PC1 and negative loading with PC2. Large (either positive or
negative) loading indicate that a variable has a strong effect on that
principal component. The larger value of cyl indicates the strong effect
on PC1.
Get biplot of these two component loadings
biplot(mtcars.pca, col = c("blue", "black"), cex = c(0.5, 1.3))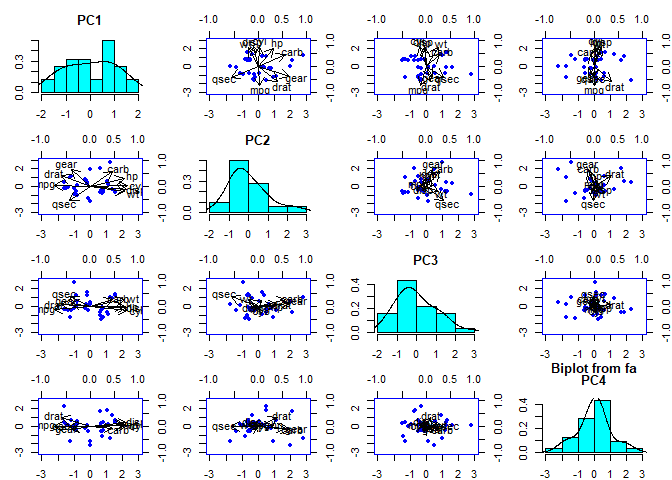
Get the head of the saved scores of mtcars.pca
head(mtcars.pca$scores)## PC1 PC2 PC3 PC4
## Mazda RX4 -0.27929417 0.8132290 -0.2877380 -0.2447853
## Mazda RX4 Wag -0.26793045 0.6770476 0.1560073 -0.1664252
## Datsun 710 -0.96699807 -0.2263347 -0.2959516 -0.2110014
## Hornet 4 Drive -0.09052843 -1.3699797 -0.4639869 -0.5984019
## Hornet Sportabout 0.66729435 -0.5743299 -1.4547534 0.2862895
## Valiant 0.02085807 -1.6956141 0.1574155 -1.6929588The original ddataset is projected into four principal components.
Get the head of the scores of first two components of mtcars.pca
head(mtcars.pca_2$scores)## PC1 PC2
## Mazda RX4 -0.27929417 0.8132290
## Mazda RX4 Wag -0.26793045 0.6770476
## Datsun 710 -0.96699807 -0.2263347
## Hornet 4 Drive -0.09052843 -1.3699797
## Hornet Sportabout 0.66729435 -0.5743299
## Valiant 0.02085807 -1.6956141Get biplot of these two component scores
biplot(mtcars.pca_2, col = c("blue", "red"))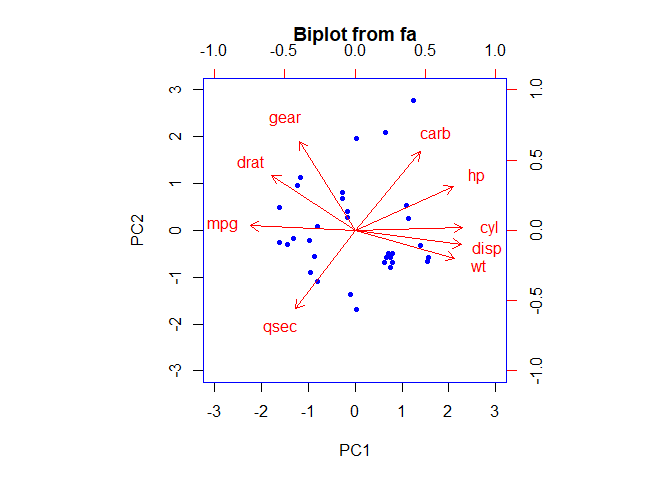
Here we can observe that hp, cyl, disp and wt contribute to PC1 with
higher values. And mpg which has negative loadings is in opposite
direction to PC1 with higher values. Gear and carb has higher
contribution to PC2 with positive values and qsec has negative value.
Get dissimilar distance of all the variables of mtcars data as mtcars.dist
#Distance calculation
mtcars.dist<- dist(mtcars.subset)Fit classical multi-dimensional scaling model with the mtcars.dist in 2-dimensional state as cars.mds.2d
# Fitting classical two- dimensional scaling model
cars.mds.2d<-cmdscale(mtcars.dist)
summary(cars.mds.2d)## V1 V2
## Min. :-181.07 Min. :-139.047
## 1st Qu.:-116.69 1st Qu.: -10.373
## Median : -43.99 Median : 2.144
## Mean : 0.00 Mean : 0.000
## 3rd Qu.: 132.85 3rd Qu.: 29.375
## Max. : 242.81 Max. : 52.503Plot the cars.mds.2d and compare it with the biplot of mtcars.pca
plot(cars.mds.2d, pch = 19)
abline(h = 0, v = 0, lty =2)
mtcars.subset<-mtcars[, 1:2] %>% scale
text(cars.mds.2d, pos = 4, labels = rownames(mtcars.subset), col = "tomato")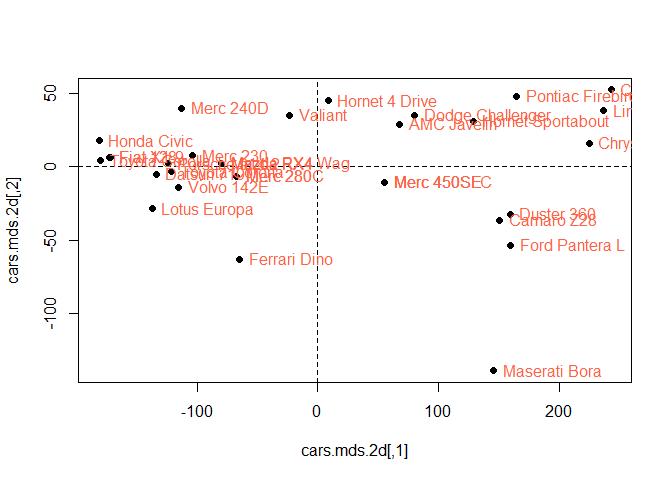
Hornet 4 Drive, Pontiac Fire bird etc lies on the positive orthant which
means they have positive contribution to the first and second
components. However, Lotus Europa and Ferari has opposite but highest
weight component 1 and component 2.
Fit classical multi-dimensional scaling model with the mtcars.dist in 3-dimensional state as cars.mds.3d
#Fiting multi-dimensional scaling model with mtcars.dist in 3 - dimensional state
cars.mds.3d<-cmdscale(mtcars.dist, k = 3)
summary(cars.mds.3d)## V1 V2 V3
## Min. :-181.07 Min. :-139.047 Min. :-6.8611
## 1st Qu.:-116.69 1st Qu.: -10.373 1st Qu.:-1.8374
## Median : -43.99 Median : 2.144 Median : 0.8492
## Mean : 0.00 Mean : 0.000 Mean : 0.0000
## 3rd Qu.: 132.85 3rd Qu.: 29.375 3rd Qu.: 2.2806
## Max. : 242.81 Max. : 52.503 Max. : 5.0029Create a 3-d scatterplot of cars.mds.3d with type = “h”, pch=20 and lty.hplot=2 and interpret it carefully
library(scatterplot3d)
cars.mds.3d <- data.frame(cmdscale(mtcars.dist, k = 3))
scatterplot3d(cars.mds.3d, type = "h", pch = 19, lty.hplot = 2)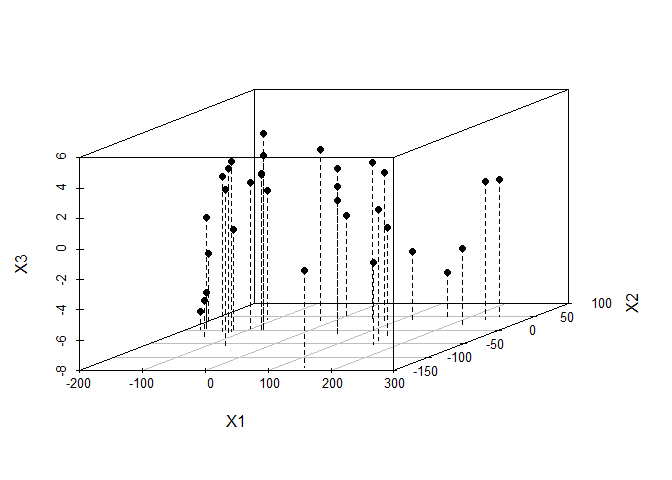
Create a 3-d scatterplot of cars.mds.3d with type = “h”, pch=20, lty.hplot=2 and color=mtcars$cyl
library(scatterplot3d)
cars.mds.3d <- data.frame(cmdscale(mtcars.dist, k = 3))
scatterplot3d(cars.mds.3d, type = "h", pch = 19, lty.hplot = 2, color = mtcars$cyl)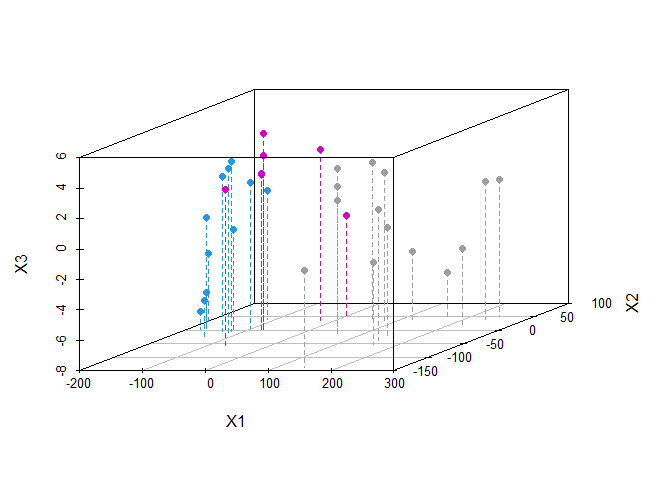
We plotted the principal components in 3- dimensional scatter plot which is
distinguished by color cyl. We can use higher dimensions by changing the
k argument in the cmdscale() function to a higher value for eg. k = 3
for 3 dimension.
Write a summary comparing PCA and MDS fits done above for mtcars data
The input to PCA is the original vectors in n-dimensional
space. Similarly, input to MDS is the pairwise distances between points.
PCA behaves as an algorithm but MDS is a visualization technique for any
factor analysis. MDS applies PCA for the dimensionality reduction. For
the mtcars, it shows that two or more but less than or equal to 5 latent
features can be generated from the given dataset.