K Medoids Clustering is a clustering algorithm, have you tried to write it from scratch in Python?
But before that, if you are also looking for other algorithms from scratch then please follow along:
- Linear Regression from Scratch
- Logistic Regression from Scratch
- Logistic Regression with Different Loss Functions
- PCA from Scratch
- K-Means Clustering from Scratch
- Feed Forward Neural Network from Scratch
- Convolutional Neural Network from Scratch
- Image Processing Class from Scratch
- Machine Learning Optimizer from Scratch
- Naive Bayes from Scratch
- Auto-Encoder from Scratch
- Basics of Stock Backtesting
K-medoids are a prominent clustering algorithm as an improvement of the predecessor, the K-Means algorithm. Despite its being widely used and less sensitive to noises and outliers, the performance of the K-medoids clustering algorithm is affected by the distance function. From here.
When the k-means algorithm is not appropriate to make objects of the cluster to the data points then the k-medoid clustering algorithm is preferred. A medoid is an object of the cluster whose dissimilarity to all the objects in the cluster is minimum. The main difference between the K-means and the K-medoid algorithm is that we work with an arbitrary matrix of distance instead of Euclidean distance. K-medoid is a classical partitioning technique of clustering that cluster the dataset into a k cluster. It is more robust to noise and outliers because it may minimize the sum of pair-wise dissimilarities however k-means minimize the sum of squared Euclidean distances. The most common distances used in KMedoids clustering techniques are Manhattan distance or Minkowski distance and here we will use Manhattan distance.
Besides K Medoids Clustering from scratch, we have also written a blog about k-means clustering from scratch. Please read it below:
Manhattan Distance
Of p1, p2 is: $|(x2-x1)+(y2-y1)|$.
Algorithm
- Step 1: Randomly select(without replacement) k of the n data points as the median.
- Step 2: Associate each data point to the closest median.
-
Step 3: While the cost of the configuration decreases:
- For each medoid
m, for each non-medoid data pointo:- Swap
mando, re-compute the cost. - If the total cost of the configuration increased in the previous step, undo the swap.
- Swap
Example
Cluster the following data set of ten objects into two clusters i.e. k =3. Data points are ${(1,3),(4,5),(6,3),(3,4),(2,1)}$
Solution
- For each medoid
Let $p_1=(1,3)$, $p_2 =(4,5)$, $p_3 =(6,3)$, $p_4=(3,4)$, $p_5 =(2,1)$. We are going to cluster the data into two clusters using the k-medoids algorithm.
Initial Step
Let $m_1= p_1=(1,3)$ and $m_2 = p_4 = (3,4)$ are two initial medoid
$ d(m_1, p_2) = mode(x_2-x_1) + mode(y_2-y_1)$
Iteration 1
Calculate the distance between cluster centers and each data point.
$d(m_1,p_1) = 0$
$d(m_2,p_1) = 3$
$d(m_1,p_2) = 3 + 2 = 5$
$d(m_2,p_2) = 1 + 1= 2$
$d(m_1,p_3) = 5 + 0 = 5$
$d(m_2,p_3) = 3 + 1 = 4$
$d(m_1,p_5) = 1 + 2= 3$
$d(m_2,p_5) = 1 + 3= 4$
Hence, after clusters are formed are
- Cluster 1 = {$p_1, p_5$}
-
Cluster 2 = {$P_2,p_3,p_4$}
Now, Total Cost = ${d(m_1,p_1) +d(m_1,p_5) + d(m_2, p_2) + d(m_2,p_3)+d(m_2,p_4)}$ = {0 + 2 + 4+3} = 9
Iteration 2Now $m_1 = p_2=(4,5) $ and $m_2 = p_4 =(3,4) $ are two medoid
Let $p_1 =(1,3)$, $p_2 =(4,5)$, $p_3=(6,3)$ and $p_5=(2,1)$.
Calculate the distance between new clusters and each data point.
$d(m_1, p_1) = 3 + 2= 5$
$d(m_2, p_1) = 2 + 1= 3$
$d(m_1, p_3) = 2 + 2 = 4$
$d(m_2, p_3) = 3 + 1 = 4$
$d(m_1, p_5) = 2 + 4 = 6 $
$d(m_2, p_5) = 1 + 3= 5$
Thus after the second iteration
- Cluster 1 = {$p_2, p_3$}
- Cluster 2 = {$p_1, p_4, p_5$}
Total cost = ${d(m_1,p_2)+d(m_1,p_3) + d(m_2, p_1) + d(m_2,p_4)+d(m_2,p_5)}$ = 0 + 4 + 3 + 0 + 5 = 12
Total cost = 12.
Here total cost in iteration 2 is greater than in iteration 1 so, we don't go further.
Coding Part
import numpy as np
import matplotlib.pyplot as plt
plt.style.use("seaborn-whitegrid")
Basic Example
Let's take an example of data points where we have 5 points and we will try to make 2 clusters for those data.
- Initially, we will plot our data points by passing
xandycoordinates of all points. - Take points 1st and 4th as initial medoids points or medoids of two clusters.
data = [(1,3),(4,5),(6,3),(3,4),(2,1)]
points = np.array(data)
# print(points)
plt.figure(figsize=(5,5))
plt.scatter(x=points[:,0],y = points[:,1])
plt.title("Initially")
plt.show()
k = 2
c1 = points[0]
c2 = points[1]
plt.figure(figsize=(5,5))
plt.scatter(x=points[:,0],y = points[:,1])
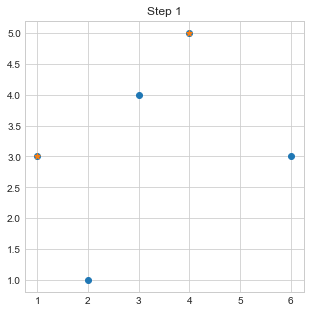
plt.scatter([c1[0],c2[0]],[c1[1],c2[1]],marker="*")
plt.title("Step 1")
plt.show()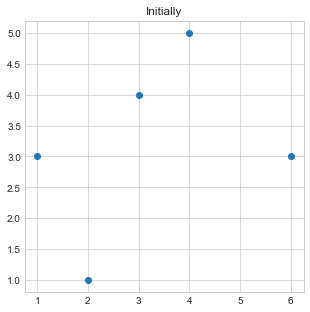
- Make a function
manhattanandget_coststo calculate the manhattan distance between points and a medoid of cluster and the minimum cost between particular data paints and medoids. - Get initial clusters and cost.
- Prepare colors for each cluster and show initial clusters.
- Now, we will loop until some internal break happens:
- Initially assume that there is no swap happened because if it is the case we must exit from the loop.
- Loop through numbers of sample
- If the current sample is not in medoid, then we will compare this sample's distance with each of the medoids.
- If We will swap the sample and medoids and calculate its cost. If the cost is smaller than the previous cost then we accept the swap and make it true as well as change the clusters. We will also plot the clusters.
- Else we ignore the swap and move on from there.
- If no swap has happened or we reached the number of iterations, then we exit from the loop.
def manhattan(p1, p2):
return np.abs((p1[0]-p2[0])) + np.abs((p1[1]-p2[1]))
def get_costs(data, medoids):
clusters = {i:[] for i in range(len(medoids))}
cst = 0
for d in data:
dst = np.array([manhattan(d, md) for md in medoids])
c = dst.argmin()
clusters[c].append(d)
cst+=dst.min()
clusters = {k:np.array(v) for k,v in clusters.items()}
return clusters, cst
def KMedoids(data, k, iters = 100):
medoids = np.array([data[i] for i in range(k)])
samples,_ = data.shape
clusters, cost = get_costs(data, medoids)
count = 0
colors = np.array(np.random.randint(0, 255, size =(k, 4)))/255
colors[:,3]=1
plt.title(f"Step : 0")
[plt.scatter(clusters[t][:, 0], clusters[t][:, 1], marker="*", s=100,
color = colors[t]) for t in range(k)]
plt.scatter(medoids[:, 0], medoids[:, 1], s=200, color=colors)
plt.show()
while True:
swap = False
for i in range(samples):
if not i in medoids:
for j in range(k):
tmp_meds = medoids.copy()
tmp_meds[j] = i
clusters_, cost_ = get_costs(data, tmp_meds)
if cost_<cost:
medoids = tmp_meds
cost = cost_
swap = True
clusters = clusters_
print(f"Medoids Changed to: {medoids}.")
plt.title(f"Step : {count+1}")
[plt.scatter(clusters[t][:, 0], clusters[t][:, 1], marker="*", s=100,
color = colors[t]) for t in range(k)]
plt.scatter(medoids[:, 0], medoids[:, 1], s=200, color=colors)
plt.show()
count+=1
if count>=iters:
print("End of the iterations.")
break
if not swap:
print("No changes.")
break
KMedoids(points,2) 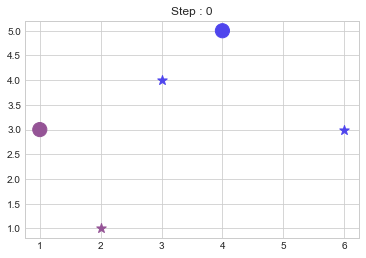
No changes.Now looking over the output of our code, it seems that it worked pretty well but what if our cluster size is variable and data also varies? So to make our code a little bit more awesome, we will do a few more tasks.
KMedoids Class
Some of the reasons that we are using a class are:
- To make our code dynamic.
- To wrap useful methods.
- To make it reusable without changing a structure.
Let's take it into action.
Initializing
- Initialize the class by giving data, the number of clusters we want, and a number of items to loop.
- Make attributes of those values.
- Take k number of medoids serially for the sake of simplicity.
- Prepare random colors for each cluster.
class KMedoidsClass:
def __init__(self,data,k,iters):
self.data= data
self.k = k
self.iters = iters
self.medoids = np.array([data[i] for i in range(self.k)])
self.colors = np.array(np.random.randint(0, 255, size =(self.k, 4)))/255
self.colors[:,3]=1Distance method
To find the distance between data points and medoids.
def manhattan(self,p1, p2):
return np.abs((p1[0]-p2[0])) + np.abs((p1[1]-p2[1]))get_cost Method
Just to find the cost minimum cost between the data points and medoids.
def get_costs(self, medoids):
tmp_clusters = {i:[] for i in range(len(medoids))}
cst = 0
for d in self.data:
dst = np.array([self.manhattan(d, md) for md in medoids])
c = dst.argmin()
tmp_clusters[c].append(d)
cst+=dst.min()
tmp_clusters = {k:np.array(v) for k,v in tmp_clusters.items()}
return tmp_clusters, cst
fit Method
This method will implement everything we did in an above simple example.
def fit(self):
samples,_ = self.data.shape
self.clusters, cost = self.get_costs(data=self.data, medoids=self.medoids)
count = 0
colors = np.array(np.random.randint(0, 255, size =(self.k, 4)))/255
colors[:,3]=1
plt.title(f"Step : 0")
[plt.scatter(self.clusters[t][:, 0], self.clusters[t][:, 1], marker="*", s=100,
color = colors[t]) for t in range(self.k)]
plt.scatter(self.medoids[:, 0], self.medoids[:, 1], s=200, color=colors)
plt.show()
while True:
swap = False
for i in range(samples):
if not i in self.medoids:
for j in range(self.k):
tmp_meds = self.medoids.copy()
tmp_meds[j] = i
clusters_, cost_ = self.get_costs(data=self.data, medoids=tmp_meds)
if cost_<cost:
self.medoids = tmp_meds
cost = cost_
swap = True
self.clusters = clusters_
print(f"Medoids Changed to: {self.medoids}.")
plt.title(f"Step : {count+1}")
[plt.scatter(self.clusters[t][:, 0], self.clusters[t][:, 1], marker="*", s=100,
color = colors[t]) for t in range(self.k)]
plt.scatter(self.medoids[:, 0], self.medoids[:, 1], s=200, color=colors)
plt.show()
count+=1
if count>=self.iters:
print("End of the iterations.")
break
if not swap:
print("No changes.")
break
Combine all code
class KMedoidsClass:
def __init__(self,data,k,iters):
self.data= data
self.k = k
self.iters = iters
self.medoids = np.array([data[i] for i in range(self.k)])
self.colors = np.array(np.random.randint(0, 255, size =(self.k, 4)))/255
self.colors[:,3]=1
def manhattan(self,p1, p2):
return np.abs((p1[0]-p2[0])) + np.abs((p1[1]-p2[1]))
def get_costs(self, medoids, data):
tmp_clusters = {i:[] for i in range(len(medoids))}
cst = 0
for d in data:
dst = np.array([self.manhattan(d, md) for md in medoids])
c = dst.argmin()
tmp_clusters[c].append(d)
cst+=dst.min()
tmp_clusters = {k:np.array(v) for k,v in tmp_clusters.items()}
return tmp_clusters, cst
def fit(self):
samples,_ = self.data.shape
self.clusters, cost = self.get_costs(data=self.data, medoids=self.medoids)
count = 0
colors = np.array(np.random.randint(0, 255, size =(self.k, 4)))/255
colors[:,3]=1
plt.title(f"Step : 0")
[plt.scatter(self.clusters[t][:, 0], self.clusters[t][:, 1], marker="*", s=100,
color = colors[t]) for t in range(self.k)]
plt.scatter(self.medoids[:, 0], self.medoids[:, 1], s=200, color=colors)
plt.show()
while True:
swap = False
for i in range(samples):
if not i in self.medoids:
for j in range(self.k):
tmp_meds = self.medoids.copy()
tmp_meds[j] = i
clusters_, cost_ = self.get_costs(data=self.data, medoids=tmp_meds)
if cost_<cost:
self.medoids = tmp_meds
cost = cost_
swap = True
self.clusters = clusters_
print(f"Medoids Changed to: {self.medoids}.")
plt.title(f"Step : {count+1}")
[plt.scatter(self.clusters[t][:, 0], self.clusters[t][:, 1], marker="*", s=100,
color = colors[t]) for t in range(self.k)]
plt.scatter(self.medoids[:, 0], self.medoids[:, 1], s=200, color=colors)
plt.show()
count+=1
if count>=self.iters:
print("End of the iterations.")
break
if not swap:
print("No changes.")
break
Test it
dt = np.random.randint(0,100, (100,2))
kmedoid = KMedoidsClass(dt,5,5)
kmedoid.fit()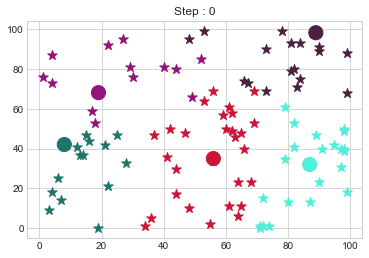
Medoids Changed to: [[10 10]
[19 68]
[56 35]
[87 32]
[89 98]].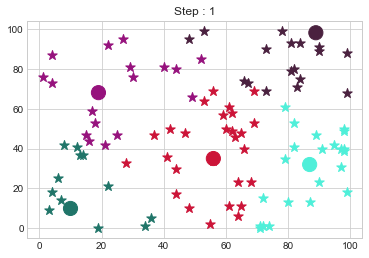
Medoids Changed to: [[11 11]
[19 68]
[56 35]
[87 32]
[89 98]].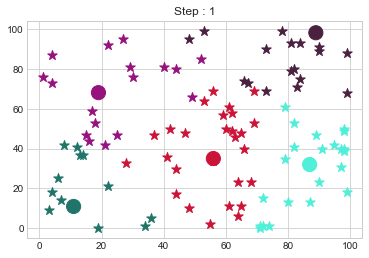
Medoids Changed to: [[12 12]
[19 68]
[56 35]
[87 32]
[89 98]].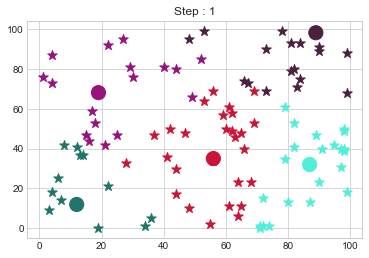
Medoids Changed to: [[13 13]
[19 68]
[56 35]
[87 32]
[89 98]].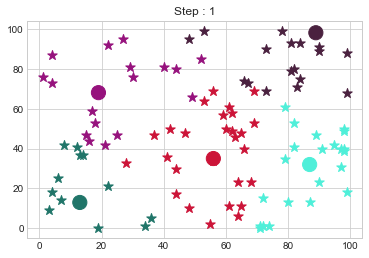
Medoids Changed to: [[14 14]
[19 68]
[56 35]
[87 32]
[89 98]].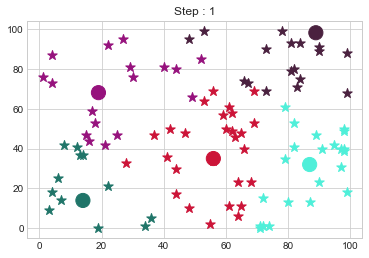
Medoids Changed to: [[18 18]
[19 68]
[56 35]
[87 32]
[89 98]].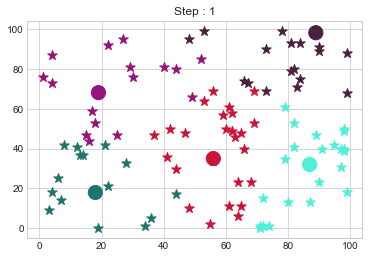
Medoids Changed to: [[18 18]
[19 68]
[49 49]
[87 32]
[89 98]].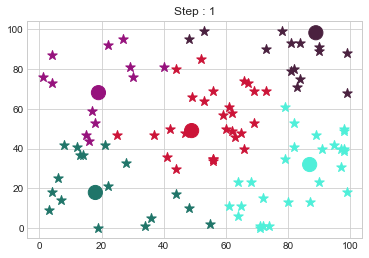
Medoids Changed to: [[18 18]
[19 68]
[50 50]
[87 32]
[89 98]].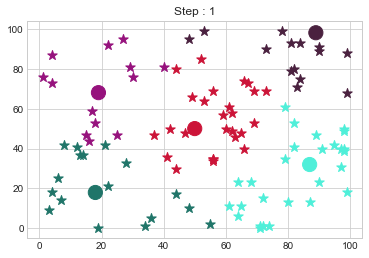
Medoids Changed to: [[18 18]
[19 68]
[51 51]
[87 32]
[89 98]].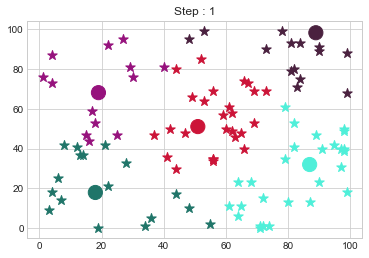
Medoids Changed to: [[18 18]
[19 68]
[52 52]
[87 32]
[89 98]].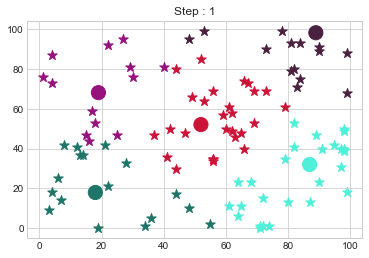
Medoids Changed to: [[18 18]
[19 68]
[53 53]
[87 32]
[89 98]].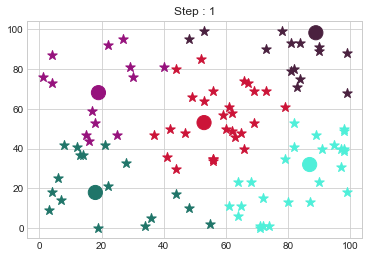
Medoids Changed to: [[18 18]
[19 68]
[54 54]
[87 32]
[89 98]].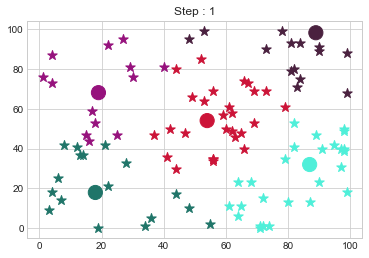
Medoids Changed to: [[18 18]
[19 68]
[55 55]
[87 32]
[89 98]].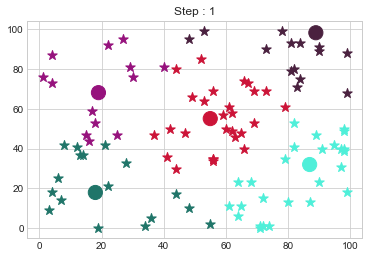
Medoids Changed to: [[18 18]
[19 68]
[56 56]
[87 32]
[89 98]].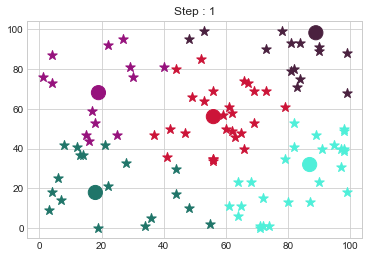
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[89 98]].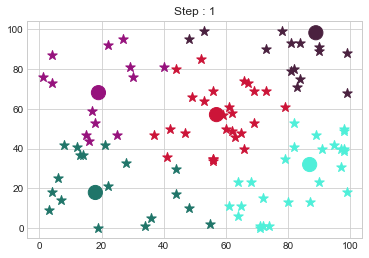
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[75 75]].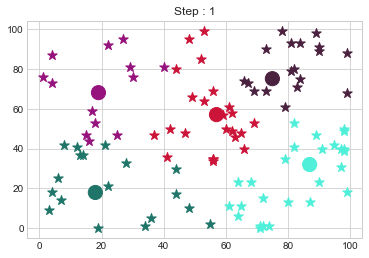
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[76 76]].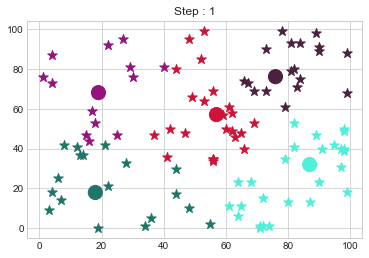
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[77 77]].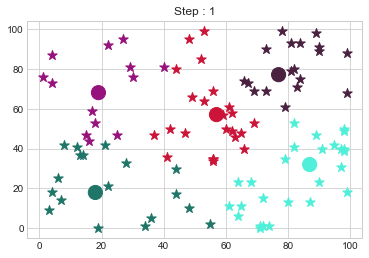
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[78 78]].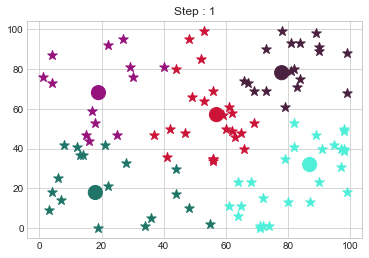
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[79 79]].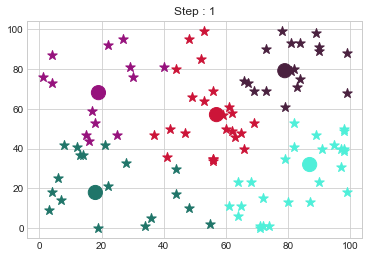
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[80 80]].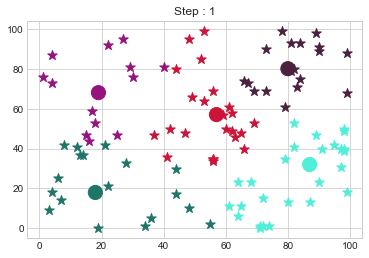
Medoids Changed to: [[18 18]
[19 68]
[57 57]
[87 32]
[81 81]].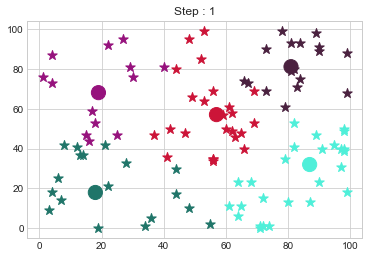
Medoids Changed to: [[20 20]
[19 68]
[57 57]
[87 32]
[81 81]].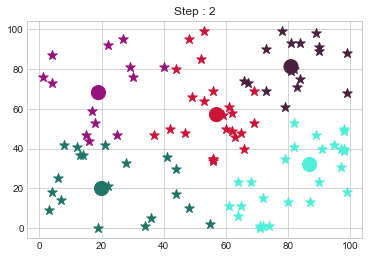
Medoids Changed to: [[21 21]
[19 68]
[57 57]
[87 32]
[81 81]].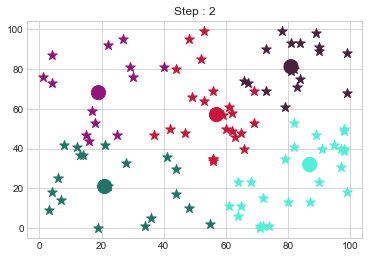
Medoids Changed to: [[21 21]
[19 68]
[51 51]
[87 32]
[81 81]].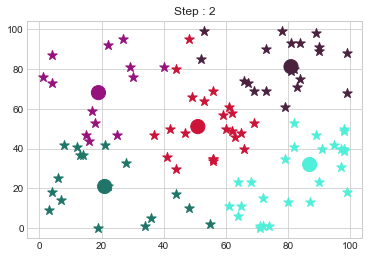
Medoids Changed to: [[21 21]
[19 68]
[52 52]
[87 32]
[81 81]].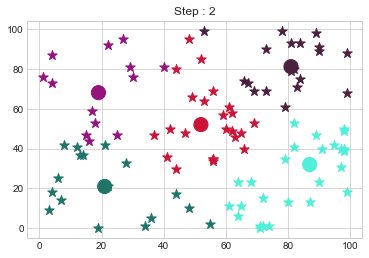
Medoids Changed to: [[21 21]
[19 68]
[53 53]
[87 32]
[81 81]].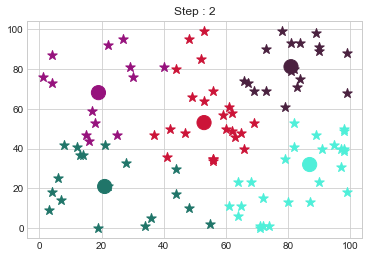
Medoids Changed to: [[16 16]
[19 68]
[53 53]
[87 32]
[81 81]].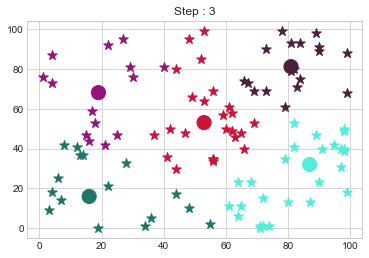
Medoids Changed to: [[17 17]
[19 68]
[53 53]
[87 32]
[81 81]].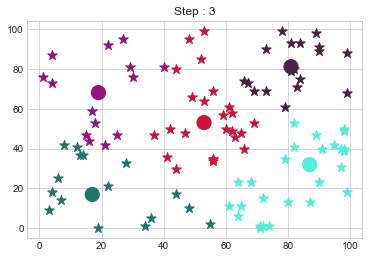
Medoids Changed to: [[18 18]
[19 68]
[53 53]
[87 32]
[81 81]].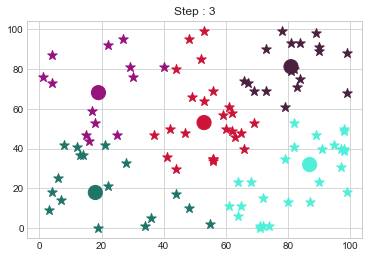
No changes.This ends the K Medoids Clustering from the the scratch blog. Thank you so much for reading this blog. In the next blog, we will do Hierarchical Clustering from Scratch.